The graph information of SGI1-B components from KU987430 |
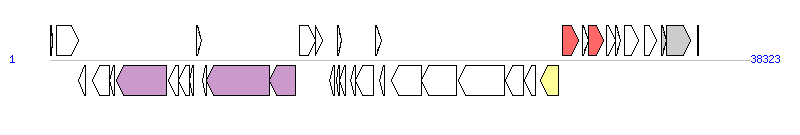 |
| Complete gene list of SGI1-B from KU987430 |
| # | Gene | Coordinates [+/-], size (bp) | Product | *Reannotation  |
| 1 | thdF | 1..163 [+], 163 | tRNA modification GTPase | |
| 2 | int | 367..1584 [+], 1218 | integrase | |
| 3 | xis | 1581..1949 [-], 369 | excisionase | |
| 4 | rep | 2307..3260 [-], 954 | replication protein | |
| 5 | S004 | 3247..3537 [-], 291 | hypothetical protein | |
| 6 | - | 3625..6384 [-], 2760 | putative mating pair stabilization protein | TraN_F, T4SS component |
| 7 | - | 6481..7014 [-], 534 | putative regulator protein | |
| 8 | - | 7017..7628 [-], 612 | hypothetical protein | |
| 9 | - | 7628..7849 [-], 222 | hypothetical protein | |
| 10 | - | 8024..8314 [+], 291 | hypothetical protein | |
| 11 | - | 8334..8588 [-], 255 | hypothetical protein | |
| 12 | - | 8588..11992 [-], 3405 | putative pilus assembly protein | TraG_F, T4SS component |
| 13 | - | 11996..13420 [-], 1425 | putative pilus assembly protein | TraH_F, T4SS component |
| 14 | - | 13664..14521 [+], 858 | hypothetical protein | |
| 15 | - | 14518..14913 [+], 396 | hypothetical protein | |
| 16 | - | 15328..15597 [-], 270 | hypothetical protein | |
| 17 | - | 15675..15860 [-], 186 | hypothetical protein | |
| 18 | - | 15720..15974 [+], 255 | hypothetical protein | |
| 19 | - | 15871..16188 [-], 318 | hypothetical protein | |
| 20 | - | 16446..16742 [-], 297 | hypothetical protein | |
| 21 | - | 16746..17711 [-], 966 | putative integrase | |
| 22 | - | 17804..18139 [+], 336 | hypothetical protein | |
| 23 | - | 18051..18326 [-], 276 | hypothetical protein | |
| 24 | - | 18687..20327 [-], 1641 | putative helicase | |
| 25 | - | 20345..22270 [-], 1926 | putative exonuclease | |
| 26 | - | 22348..24891 [-], 2544 | putative subtilisin proteinase-like protein | |
| 27 | - | 24912..25901 [-], 990 | putative ATPase | |
| 28 | res | 25980..26564 [-], 585 | resolvase | |
| 29 | intI1 | 26838..27851 [-], 1014 | IntI1 integrase | Integrase |
| 30 | blaPSE-1 | 28057..28923 [+], 867 | beta-lactamase | AR |
| 31 | qacEdelta1 | 29140..29487 [+], 348 | QacEdelta1 | |
| 32 | sul1 | 29481..30320 [+], 840 | dihydropteroate synthase | AR |
| 33 | - | 30448..30948 [+], 501 | acetyltransferase | |
| 34 | - | 30972..31259 [+], 288 | hypothetical protein | |
| 35 | - | 31425..32219 [+], 795 | IS6100 transposase | |
| 36 | S044 | 32567..33196 [+], 630 | hypothetical protein | |
| 37 | hipB | 33472..33729 [+], 258 | transcriptional regulator | |
| 38 | hipA | 33730..35043 [+], 1314 | regulatory protein | |
| 39 | PMI3124 | 35446..35500 [+], 55 | membrane protein | |
