The graph information of MTnPi4 components from AP014597 |
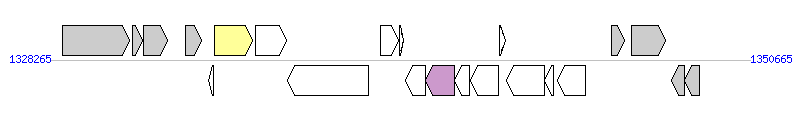 |
| Complete gene list of MTnPi4 from AP014597 |
| # | Gene | Coordinates [+/-], size (bp) | Product | *Reannotation  |
| 1 | PIOMA14_I_1205 | 1328650..1330809 [+], 2160 | conserved hypothetical protein | |
| 2 | PIOMA14_I_1206 | 1330915..1331235 [+], 321 | conserved hypothetical protein with DUF59 domain | |
| 3 | PIOMA14_I_1207 | 1331270..1332034 [+], 765 | UDP-2,3-diacylglucosamine hydrolase | |
| 4 | PIOMA14_I_1208 | 1332591..1333115 [+], 525 | putative DNA mismatch endonuclease Vsr | |
| 5 | PIOMA14_I_1209 | 1333329..1333499 [-], 171 | hypothetical protein | |
| 6 | PIOMA14_I_1210 | 1333521..1334744 [+], 1224 | integrase | Integrase |
| 7 | PIOMA14_I_1211 | 1334852..1335841 [+], 990 | probable ATPase | |
| 8 | PIOMA14_I_1212 | 1335865..1338471 [-], 2607 | probable peptidase subtilase family | |
| 9 | PIOMA14_I_1213 | 1338838..1339428 [+], 591 | conserved hypothetical protein with HATPase-c domain | |
| 10 | PIOMA14_I_1214 | 1339454..1339573 [+], 120 | hypothetical protein | |
| 11 | PIOMA14_I_1215 | 1339632..1340279 [-], 648 | conserved hypothetical protein | |
| 12 | PIOMA14_I_1216 | 1340291..1341211 [-], 921 | probable relaxase | Relaxase |
| 13 | PIOMA14_I_1217 | 1341208..1341702 [-], 495 | hypothetical protein | |
| 14 | PIOMA14_I_1218 | 1341705..1342616 [-], 912 | DNA primase | |
| 15 | PIOMA14_I_1219 | 1342657..1342830 [+], 174 | hypothetical protein | |
| 16 | PIOMA14_I_1220 | 1342884..1344074 [-], 1191 | conserved hypothetical protein | |
| 17 | PIOMA14_I_1221 | 1344078..1344371 [-], 294 | probable excisionase | |
| 18 | PIOMA14_I_1222 | 1344496..1345416 [-], 921 | conserved hypothetical protein | |
| 19 | PIOMA14_I_1223 | 1346231..1346656 [+], 426 | hypothetical protein with DUF2147 domain | |
| 20 | PIOMA14_I_1224 | 1346874..1347968 [+], 1095 | mannose-1-phosphate guanylyltransferase | |
| 21 | PIOMA14_I_1225 | 1348159..1348557 [-], 399 | probable histidine triad (HIT) family protein | |
| 22 | PIOMA14_I_1226 | 1348582..1349052 [-], 471 | putative transcription elongation factor GreA | |
