The graph information of MGIPspArc1 components from NZ_BADU01000066 |
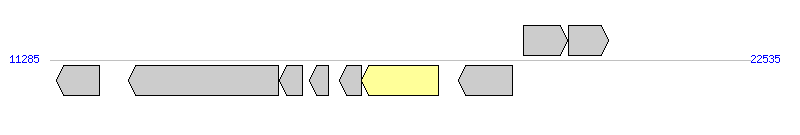 |
| Incomplete gene list of MGIPspArc1 from NZ_BADU01000066 |
| # | Gene | Coordinates [+/-], size (bp) | Product | *Reannotation  |
| 1 | P20311_RS07340 | 11389..12075 [-], 687 | hypothetical protein | |
| 2 | P20311_RS07345 | 12543..14951 [-], 2409 | DNA helicase | |
| 3 | P20311_RS07350 | 14977..15348 [-], 372 | transcriptional regulator | |
| 4 | P20311_RS07355 | 15448..15753 [-], 306 | type II toxin-antitoxin system HigB family toxin | |
| 5 | P20311_RS07360 | 15945..16283 [-], 339 | hypothetical protein | |
| 6 | P20311_RS07365 | 16285..17535 [-], 1251 | site-specific integrase | Integrase |
| 7 | P20311_RS07370 | 17851..18711 [-], 861 | YicC family protein | |
| 8 | P20311_RS07375 | 18892..19605 [+], 714 | ribonuclease PH | |
| 9 | P20311_RS07380 | 19620..20264 [+], 645 | orotate phosphoribosyltransferase | |
