The graph information of ICEBs1 components from AL009126 |
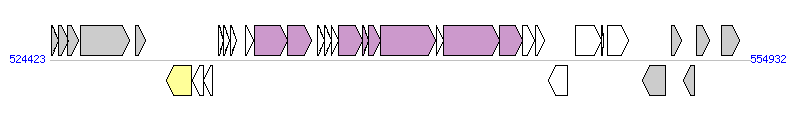 |
| Complete gene list of ICEBs1 from AL009126 |
| # | Gene  | Coordinates [+/-], size (bp) | Product
(GenBank annotation) | *Reannotation  |
| 1 | ydcF | 524492..524785 [+], 294 | hypothetical protein | |
| 2 | ydcG | 524782..525222 [+], 441 | conserved hypothetical protein | |
| 3 | ydcH | 525206..525649 [+], 444 | putative transcriptional regulator | |
| 4 | ydcI | 525743..527902 [+], 2160 | putative RNA helicase | |
| 5 | ydcK | 528129..528581 [+], 453 | conserved hypothetical protein | |
| 6 | ydcL (int) | 529505..530611 [-], 1107 | putative phage integrase | Integrase |
| 7 | immA | 530624..531133 [-], 510 | immunity anti-repressor conserved in prophages | |
| 8 | immR | 531130..531513 [-], 384 | phage element (ICEBs1)transcriptional regulator (Xre family) | |
| 9 | sacV (xis) | 531787..531981 [+], 195 | transcriptional regulator | |
| 10 | ydzL | 531978..532238 [+], 261 | conserved hypothetical protein | |
| 11 | ydcO | 532292..532552 [+], 261 | hypothetical protein; mobile element region | |
| 12 | ydcP (helP) | 532922..533302 [+], 381 | conserved hypothetical protein; mobile element region | HelP; ICEBs1-encoded helicase processivity factor; [PMID: 23326247] |
| 13 | ydcQ (conQ) | 533338..534780 [+], 1443 | putative DNA wielding protein; mobile element region | YdcQ, T4SS component |
| 14 | ydcR (nicK) | 534773..535831 [+], 1059 | putative replication protein; mobile element region | Relaxase, MOBT Family |
| 15 | ydcS | 536096..536365 [+], 270 | conserved hypothetical protein; mobile element region | |
| 16 | ydcT | 536404..536670 [+], 267 | conserved hypothetical protein; mobile element region | |
| 17 | yddA | 536687..536995 [+], 309 | conserved hypothetical protein; mobile element region | |
| 18 | yddB (conB) | 536985..538049 [+], 1065 | conserved hypothetical protein; mobile element region | YddB, T4SS component |
| 19 | yddC (conC) | 538061..538309 [+], 249 | conserved hypothetical protein | YddC, T4SS component |
| 20 | yddD (conD) | 538322..538846 [+], 525 | hypothetical protein; mobile element region | YddD, T4SS component |
| 21 | yddE (conE) | 538836..541229 [+], 2394 | conserved hypothetical protein | YddE, T4SS component |
| 22 | yddF | 541248..541574 [+], 327 | conserved hypothetical protein; mobile element region | |
| 23 | yddG (conG) | 541578..544025 [+], 2448 | conserved hypothetical protein; mobile element region | YddG, T4SS component |
| 24 | yddH (cwlT) | 544022..545011 [+], 990 | putative cell wall hydrolase; mobile element region | YddH, T4SS component |
| 25 | yddI | 545026..545532 [+], 507 | conserved hypothetical protein; mobile element region | |
| 26 | yddJ | 545595..545975 [+], 381 | putative lipoprotein | |
| 27 | yddK | 546166..546966 [-], 801 | conserved hypothetical protein; mobile element region | |
| 28 | rapI | 547306..548481 [+], 1176 | response regulator aspartate phosphatase; mobile element region | |
| 29 | phrI | 548438..548557 [+], 120 | secreted regulator of the activity of phosphatase RapI | |
| 30 | yddM | 548710..549651 [+], 942 | putative helicase; mobile element region | |
| 31 | yddN | 550240..551259 [-], 1020 | putative alkanal monooxygenase | |
| 32 | lrpA | 551519..551929 [+], 411 | transcriptional regulator (Lrp/AsnC family) | |
| 33 | lrpB | 552052..552501 [-], 450 | transcriptional regulator (Lrp/AsnC family) | |
| 34 | yddQ | 552616..553158 [+], 543 | putative hydrolase | |
| 35 | yddR | 553711..554475 [+], 765 | putative metal-dependent hydrolase | |

