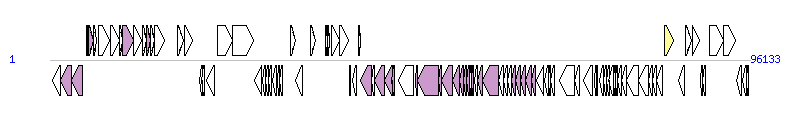The graph information of ICESe4 components from FR686852 |
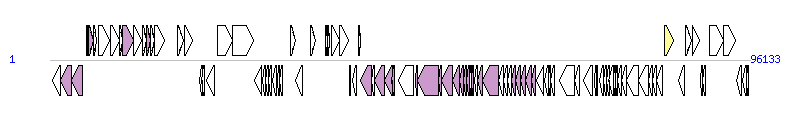 |
| Complete gene list of ICESe4 from FR686852 |
| # | Gene  | Coordinates [+/-], size (bp) | Product
(GenBank annotation) | *Reannotation  |
| 1 | SHAD27981 | 281..1381 [-], 1101 | putative integrase (phage) | |
| 2 | SHAD27991 | 1424..2971 [-], 1548 | putative helicase | Relaxase, MOBH Family |
| 3 | SHAD28001 | 2946..4454 [-], 1509 | conserved hypothetical protein | TraI_F, T4SS component |
| 4 | SHAD28020 | 4981..5199 [+], 219 | conserved hypothetical protein | |
| 5 | pilL | 5252..5977 [+], 726 | putative PilL exported protein | Tfc2, T4SS component |
| 6 | pilM | 5980..6417 [+], 438 | putative PilM exported protein | |
| 7 | pilN | 6607..8232 [+], 1626 | putative PilN pilus assembly protein | |
| 8 | pilO | 8244..9542 [+], 1299 | putative PilO pilin accessory protein | |
| 9 | pilP | 9532..9993 [+], 462 | putative PilP, pilus assembly protein | |
| 10 | pilQ | 10003..11523 [+], 1521 | Putative PilQ, nucleotide binding protein | TraJ_I, T4SS component |
| 11 | pilR | 11525..12625 [+], 1101 | putative PilR membrane protein | |
| 12 | pilS | 12675..13211 [+], 537 | putative PilS, major pilin subunit protein | |
| 13 | pilT | 13270..13746 [+], 477 | putative PilT, pilin transglycosylase | Orf169_F, T4SS component |
| 14 | pilU | 13759..14409 [+], 651 | putative pilU, prepilin peptidase | |
| 15 | pilV | 14406..15734 [+], 1329 | putative PilV, minor pilin subunit protein | |
| 16 | traE | 17489..18310 [+], 822 | putative TraE protein | |
| 17 | SHAD28161 | 18412..19614 [+], 1203 | putative TraF protein | |
| 18 | SHAD28170 | 20551..20967 [-], 417 | putative transposase | |
| 19 | SHAD28171 | 20858..21199 [-], 342 | putative transposase | |
| 20 | SHAD28181 | 21532..22545 [-], 1014 | conserved hypothetical protein | |
| 21 | SHAD28191 | 23022..25058 [+], 2037 | type III restriction endonuclease | |
| 22 | SHAD28201 | 25061..27976 [+], 2916 | type III restriction endonuclease | |
| 23 | SHAD28211 | 28058..29035 [-], 978 | conserved hypothetical protein | |
| 24 | SHAD28221 | 29143..29580 [-], 438 | Hypothetical protein | |
| 25 | SHAD28231 | 29656..29994 [-], 339 | Hypothetical protein | |
| 26 | SHAD28241 | 30062..30421 [-], 360 | Hypothetical protein | |
| 27 | SHAD28251 | 30618..31127 [-], 510 | putative antirestriction protein | |
| 28 | SHAD28252 | 31268..31459 [-], 192 | conserved hypothetical protein | |
| 29 | SHAD28261 | 31546..31932 [-], 387 | Hypothetical protein | |
| 30 | SHAD28272 | 33016..33756 [+], 741 | conserved hypothetical protein | |
| 31 | SHAD28281 | 33760..34677 [-], 918 | conserved hypothetical protein | |
| 32 | SHAD28291 | 35824..36477 [+], 654 | conserved hypothetical protein | |
| 33 | SHAD28301 | 37816..38064 [+], 249 | putative antitoxin protein (ccdA-like) | |
| 34 | SHAD28311 | 38064..38387 [+], 324 | putative toxin protein (CcdB-like) | |
| 35 | SHAD28321 | 38623..39777 [+], 1155 | putative methyltransferase | |
| 36 | SHAD28331 | 39793..40965 [+], 1173 | conserved hypothetical protein | |
| 37 | rpmE | 41072..41335 [-], 264 | ribosomal protein L31 | |
| 38 | SHAD28351 | 41559..42131 [-], 573 | conserved hypothetical protein | |
| 39 | SHAD28361 | 42429..42635 [+], 207 | conserved hypothetical protein | |
| 40 | SHAD28371 | 42683..44206 [-], 1524 | putative membrane protein | Tfc19, T4SS component |
| 41 | SHAD28381 | 44203..44562 [-], 360 | putative exported protein | |
| 42 | SHAD28391 | 44567..45985 [-], 1419 | putative exported protein | Tfc22, T4SS component |
| 43 | SHAD28401 | 45995..46969 [-], 975 | putative exported protein | Tfc23, T4SS component |
| 44 | SHAD28411 | 46966..47361 [-], 396 | putative exported protein | Tfc24, T4SS component |
| 45 | SHAD28421 | 47899..49986 [-], 2088 | putative adhesin / TonB-dependent uptake protein | |
| 46 | SHAD28431 | 50142..50528 [-], 387 | conserved hypothetical protein | Tfc17, T4SS component |
| 47 | SHAD28441 | 50525..53374 [-], 2850 | conserved hypothetical protein | Tfc16, T4SS component |
| 48 | SHAD28451 | 53374..53778 [-], 405 | putative lipoprotein precursor | Tfc15, T4SS component |
| 49 | SHAD28461 | 53796..55274 [-], 1479 | conserved hypothetical protein | Tfc14, T4SS component |
| 50 | SHAD28471 | 55264..56184 [-], 921 | conserved hypothetical protein | Tfc13, T4SS component |
| 51 | SHAD28481 | 56181..56828 [-], 648 | hypothetical protein | Tfc12, T4SS component |
| 52 | SHAD28482 | 56825..57190 [-], 366 | conserved hypothetical protein | Tfc11, T4SS component |
| 53 | SHAD28501 | 57209..57574 [-], 366 | conserved hypothetical protein | Tfc10, T4SS component |
| 54 | SHAD28502 | 57602..57844 [-], 243 | conserved hypothetical protein | |
| 55 | SHAD28521 | 57844..58185 [-], 342 | conserved hypothetical protein | Tfc9, T4SS component |
| 56 | SHAD28522 | 58365..58700 [-], 336 | hypothetical protein | |
| 57 | SHAD28531 | 58704..59465 [-], 762 | putative membrane protein | Tfc8, T4SS component |
| 58 | SHAD28541 | 59446..61563 [-], 2118 | putative TraD/TraG protein | Tfc6, T4SS component |
| 59 | SHAD28551 | 61563..62348 [-], 786 | hypothetical protein | |
| 60 | SHAD28561 | 62345..62857 [-], 513 | hypothetical protein | |
| 61 | SHAD28571 | 62854..63444 [-], 591 | putative membrane protein | |
| 62 | SHAD28581 | 63444..63971 [-], 528 | conserved hypothetical protein | Tfc5, T4SS component |
| 63 | SHAD28591 | 63968..64624 [-], 657 | putative exported protein | Tfc4, T4SS component |
| 64 | SHAD28601 | 64603..65322 [-], 720 | putative membrane protein | Tfc3, T4SS component |
| 65 | SHAD28611 | 65327..66106 [-], 780 | putative membrane protein | Tfc2, T4SS component |
| 66 | SHAD28621 | 66103..66681 [-], 579 | putative lipoprotein | Tfc2, T4SS component |
| 67 | SHAD28631 | 66872..67837 [-], 966 | putative transposase | |
| 68 | SHAD28641 | 67911..68468 [-], 558 | single-stranded DNA-binding protein | |
| 69 | SHAD28642 | 68481..68684 [-], 204 | conserved hypothetical protein | |
| 70 | SHAD28661 | 68784..69248 [-], 465 | conserved hypothetical protein | |
| 71 | topB | 69959..71977 [-], 2019 | DNA topoisomerase | |
| 72 | SHAD28681 | 71990..72670 [-], 681 | conserved hypothetical protein | |
| 73 | SHAD28701 | 73245..74510 [-], 1266 | conserved hypothetical protein | |
| 74 | SHAD28710 | 74771..74992 [-], 222 | hypothetical protein | |
| 75 | SHAD28711 | 74989..75204 [-], 216 | hypothetical protein | |
| 76 | SHAD28713 | 75447..75962 [-], 516 | conserved hypothetical | |
| 77 | SHAD28714 | 75964..76461 [-], 498 | conserved hypothetical protein | |
| 78 | SHAD28721 | 76458..76772 [-], 315 | conserved hypothetical protein | |
| 79 | SHAD28731 | 76769..77377 [-], 609 | conserved hypothetical protein | |
| 80 | SHAD28732 | 77364..77621 [-], 258 | hypothetical protein | |
| 81 | SHAD28733 | 77631..77879 [-], 249 | hypothetical protein | |
| 82 | SHAD28761 | 77876..78454 [-], 579 | conserved hypothetical protein | |
| 83 | SHAD28771 | 78451..79182 [-], 732 | conserved hypothetical protein | |
| 84 | SHAD28781 | 79175..80824 [-], 1650 | conserved hypothetical protein | |
| 85 | SHAD28791 | 80814..82202 [-], 1389 | putative DNA helicase | |
| 86 | SHAD28792 | 82195..82707 [-], 513 | hypothetical protein | |
| 87 | SHAD28793 | 82694..83257 [-], 564 | hypothetical protein | |
| 88 | SHAD28811 | 83277..84155 [-], 879 | putative parA partitioning protein | |
| 89 | SHAD28821 | 84416..85675 [+], 1260 | putative integrase (phage) | Integrase |
| 90 | SHAD28831 | 86256..87143 [-], 888 | putative DNA-binding protein (phage) | |
| 91 | SHAD28841 | 87281..88129 [+], 849 | hypothetical protein | |
| 92 | SHAD28851 | 88241..89260 [+], 1020 | putative nucleotide binding protein | |
| 93 | SHAD28861 | 89353..89592 [-], 240 | putative transcriptional regulator (phage) | |
| 94 | SHAD28871 | 89589..90050 [-], 462 | conserved hypothetical protein | |
| 95 | SHAD28881 | 90565..92478 [+], 1914 | hypothetical protein | |
| 96 | SHAD28891 | 92475..94175 [+], 1701 | conserved hypothetical protein | |
| 97 | SHAD28901 | 94318..95016 [-], 699 | hypothetical protein | |
| 98 | SHAD28911 | 95103..95594 [-], 492 | putative acetyltransferase | |
| 99 | SHAD28921 | 95591..95884 [-], 294 | conserved hypothetical protein | |