The graph information of Tn4371 components from AJ536756 |
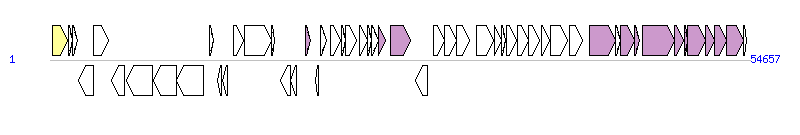 |
| Complete gene list of Tn4371 from AJ536756 |
| # | Gene  | Coordinates [+/-], size (bp) | Product
(GenBank annotation) | *Reannotation  |
| 1 | int | 181..1389 [+], 1209 | putative site specific recombinase | Integrase |
| 2 | - | 1469..1765 [+], 297 | hypothetical protein | |
| 3 | - | 1762..2169 [+], 408 | hypothetical protein | |
| 4 | - | 2210..3385 [-], 1176 | hypothetical protein | |
| 5 | - | 3388..4533 [+], 1146 | putative transposase | |
| 6 | - | 4813..5835 [-], 1023 | hypothetical protein | |
| 7 | - | 5989..7980 [-], 1992 | hypothetical protein | |
| 8 | - | 8072..9871 [-], 1800 | hypothetical protein | |
| 9 | - | 9874..12012 [-], 2139 | hypothetical protein | |
| 10 | - | 12442..12792 [+], 351 | hypothetical protein | |
| 11 | - | 13054..13410 [-], 357 | hypothetical protein | |
| 12 | - | 13502..13882 [-], 381 | hypothetical protein | |
| 13 | - | 14290..15117 [+], 828 | hypothetical protein | |
| 14 | - | 15194..17266 [+], 2073 | hypothetical protein | |
| 15 | - | 17330..17539 [+], 210 | hypothetical protein | |
| 16 | - | 18015..18785 [-], 771 | hypothetical protein | |
| 17 | - | 18782..19276 [-], 495 | putative transcription regulator | |
| 18 | - | 19931..20329 [+], 399 | hypothetical protein | TraF, T4SS component |
| 19 | - | 20697..20990 [-], 294 | putative transcription regulator | |
| 20 | - | 21160..21561 [+], 402 | hypothetical protein | |
| 21 | - | 21903..22664 [+], 762 | hypothetical protein | |
| 22 | - | 22748..23038 [+], 291 | hypothetical protein | |
| 23 | repA | 23065..23922 [+], 858 | putative replication initiation protein | |
| 24 | parA | 24188..24826 [+], 639 | putative plasmid partition protein | |
| 25 | parB | 24823..25086 [+], 264 | hypothetical protein | |
| 26 | - | 25083..25616 [+], 534 | hypothetical protein | |
| 27 | traF | 25613..26212 [+], 600 | putative plasmid conjugal transfer transmembrane protein | TraF, T4SS component |
| 28 | - | 26607..28160 [+], 1554 | hypothetical protein | Relaxase, MOBL Family |
| 29 | bphS | 28524..29468 [-], 945 | transcription regulator | |
| 30 | bphE | 29962..30768 [+], 807 | 2-oxopent-4-enoate hydratase | |
| 31 | bphG | 30771..31712 [+], 942 | acetaldehyde dehydrogenase oxidoreductase | |
| 32 | bphF | 31724..32779 [+], 1056 | putative 4-hydroxy-2-oxovalerate aldolase transmembrane protein | |
| 33 | bphA1 | 33268..34644 [+], 1377 | biphenyl 2,3-dioxygenase oxidoreductase | |
| 34 | bphA2 | 34676..35257 [+], 582 | biphenyl 2,3-dioxygenase | |
| 35 | bphA3 | 35275..35604 [+], 330 | 3-phenylpropionate dioxygenase ferredoxin oxidoreductase | |
| 36 | bphB | 35623..36453 [+], 831 | 2,3-dihydroxy-2,3-dihydro-phenylpropionate dehydrogenase | |
| 37 | bphC | 36467..37348 [+], 882 | 2,3-dihydroxybiphenyl dioxygenase | |
| 38 | bphD | 37380..38240 [+], 861 | 2-hydroxy-6-ketonona-2,4-dienedioic acid hydrolase | |
| 39 | - | 38365..39021 [+], 657 | hypothetical protein | |
| 40 | bphA4 | 39116..40411 [+], 1296 | 3-phenylpropionate dioxygenase ferredoxin oxidoreductase | |
| 41 | traR | 40563..41606 [+], 1044 | transcription regulator | |
| 42 | traG | 42118..44121 [+], 2004 | putative conjugal transfer protein | VirD4, T4SS component |
| 43 | - | 44118..44582 [+], 465 | hypothetical protein | |
| 44 | trbB | 44579..45616 [+], 1038 | putative mating pair formation protein | TrbB, T4SS component |
| 45 | trbC | 45622..46005 [+], 384 | putative mating pair formation transmembrane protein | TrbC, T4SS component |
| 46 | trbE | 46287..48740 [+], 2454 | putative mating pair formation protein | TrbE, T4SS component |
| 47 | trbJ | 48737..49492 [+], 756 | putative mating pair formation protein | TrbJ, T4SS component |
| 48 | - | 49504..49803 [+], 300 | hypothetical protein | TrbJ, T4SS component |
| 49 | trbL | 49800..51149 [+], 1350 | putative mating pair formation transmembrane protein | TrbL, T4SS component |
| 50 | trbF | 51168..51872 [+], 705 | putative mating pair formation protein | TrbF, T4SS component |
| 51 | trbG | 51869..52849 [+], 981 | putative mating pair formation protein | TrbG, T4SS component |
| 52 | trbI | 52852..54132 [+], 1281 | putative mating pair formation transmembrane protein | TrbI, T4SS component |
| 53 | - | 54129..54374 [+], 246 | hypothetical protein | |
