The graph information of ICESp1108 components from FR691054 |
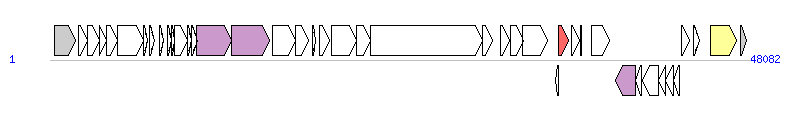 |
| Complete gene list of ICESp1108 from FR691054 |
| # | Gene  | Coordinates [+/-], size (bp) | Product
(GenBank annotation) | *Reannotation  |
| 1 | rum | 332..1720 [+], 1389 | 23S rRNA m(5)U 1939 methyltransferase | |
| 2 | ORF1 | 1951..2586 [+], 636 | replication initiator protein | |
| 3 | ORF2 | 2583..3410 [+], 828 | DNA replication protein | |
| 4 | ORF3 | 3403..3888 [+], 486 | hypothetical protein | |
| 5 | ORF4 | 3885..4619 [+], 735 | phage antirepressor protein | |
| 6 | ORF5 | 4612..6408 [+], 1797 | TraG/TraD/VirD4 bacterial conjugation protein | |
| 7 | ORF6 | 6427..6702 [+], 276 | hypothetical protein | |
| 8 | ORF7 | 6814..7182 [+], 369 | hypothetical protein | |
| 9 | ORF8 | 7529..7822 [+], 294 | hypothetical protein | |
| 10 | ORF9 | 8045..8356 [+], 312 | hypothetical protein | |
| 11 | ORF10 | 8358..8573 [+], 216 | hypothetical protein | |
| 12 | ORF11 | 8584..9447 [+], 864 | hypothetical protein | |
| 13 | ORF12 | 9457..9747 [+], 291 | hypothetical protein | |
| 14 | ORF13 | 9749..10159 [+], 411 | bacitracin resistance protein | |
| 15 | ORF14 | 10050..12470 [+], 2421 | type IV secretory pathway protein VirB4 | PrgJ, T4SS component |
| 16 | ORF15 | 12478..15057 [+], 2580 | NlpC/P60 family protein | Orf14_Tn, T4SS component |
| 17 | ORF16 | 15298..16881 [+], 1584 | amine oxidase | |
| 18 | ORF17 | 16896..17756 [+], 861 | hypothetical protein | |
| 19 | ORF18 | 18053..18256 [+], 204 | hypothetical protein | |
| 20 | ORF19 | 18492..19232 [+], 741 | hypothetical protein | |
| 21 | ORF20 | 19354..21087 [+], 1734 | DNA topoisomerase IA | |
| 22 | ORF21 | 21080..22045 [+], 966 | C-5 cytosine specific DNA methylase | |
| 23 | ORF22 | 22032..29675 [+], 7644 | superfamily II DNA/RNA helicase | |
| 24 | ORF23 | 29719..30375 [+], 657 | hypothetical protein | |
| 25 | ORF24 | 30927..31559 [+], 633 | transcriptional regulator, TetR family | |
| 26 | ORF25 | 31644..32525 [+], 882 | tetronasin resistance ATP-binding protein | |
| 27 | ORF26 | 32458..34164 [+], 1707 | tetronasin resistance transmembrane protein | |
| 28 | ORF27 | 34745..34951 [-], 207 | hypothetical protein | |
| 29 | erm(TR) | 34898..35629 [+], 732 | erythromycin resistance methylase | AR |
| 30 | ORF29 | 35846..36370 [+], 525 | spectinomycin phosphotransferase | |
| 31 | ORF30 | 36416..36535 [+], 120 | hypothetical protein | |
| 32 | ORF31 | 37228..38451 [+], 1224 | transposase IS116/IS110/IS902 family protein | |
| 33 | ORF32 | 38857..40188 [-], 1332 | relaxase/mobilization protein | Relaxase, MOBP Family |
| 34 | ORF33 | 40190..40648 [-], 459 | hypothetical protein | |
| 35 | ORF34 | 40717..41811 [-], 1095 | RhuM | |
| 36 | ORF35 | 41792..42274 [-], 483 | cation transporter family protein | |
| 37 | ORF36 | 42283..42831 [-], 549 | Zn-dependent peptidase | |
| 38 | ORF37 | 42840..43241 [-], 402 | trascriptional regulator Cro/CI family protein | |
| 39 | ORF38 | 43376..43951 [+], 576 | hypothetical protein | |
| 40 | ORF39 | 44221..44631 [+], 411 | RNA polymerase sigma-70 region 4 | |
| 41 | ORF40 | 45362..47149 [+], 1788 | putative recombinase | Integrase |
| 42 | pnp | 47452..47850 [+], 399 | phosphorylase | |
