The graph information of ICEPmiUSA1 components from AM942759 |
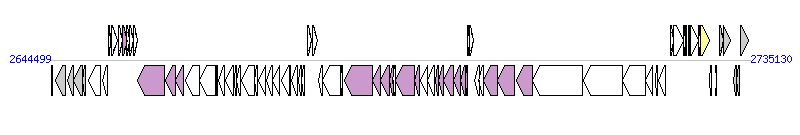 |
| Complete gene list of ICEPmiUSA1 from AM942759 |
| # | Gene  | Coordinates [+/-], size (bp) | Product
(GenBank annotation) | *Reannotation  |
| 1 | PMI2417 | 2644690..2644863 [-], 174 | hypothetical protein | |
| 2 | PMI2418 | 2645217..2646518 [-], 1302 | putative Na+ dependent nucleoside transporter | |
| 3 | PMI2419 | 2646806..2647588 [-], 783 | putative TatD-related deoxyribonuclease | |
| 4 | PMI2420 | 2647585..2648646 [-], 1062 | patatin-like phospholipase | |
| 5 | PMI2421 | 2648743..2649090 [-], 348 | putative phospholipid-binding protein | |
| 6 | prfC | 2649499..2651088 [-], 1590 | peptide chain release factor 3 | |
| 7 | PMI2423 | 2651345..2651992 [-], 648 | putative phage repressor | |
| 8 | PMI2424 | 2652110..2652361 [+], 252 | phage regulatory protein | |
| 9 | PMI2425 | 2652417..2653286 [+], 870 | putative phage protein | |
| 10 | PMI2426 | 2653327..2653935 [+], 609 | putative phage protein | |
| 11 | PMI2427 | 2653922..2654470 [+], 549 | putative phage transglycosylase | Orf169_F, T4SS component |
| 12 | PMI2428 | 2654467..2654766 [+], 300 | putative phage protein | |
| 13 | PMI2429 | 2654763..2655296 [+], 534 | putative phage regulatory protein | |
| 14 | PMI2430 | 2655354..2655785 [+], 432 | putative phage membrane protein | |
| 15 | PMI2431 | 2655818..2659387 [-], 3570 | putative pilus assembly protein | TraG_F, T4SS component |
| 16 | PMI2432 | 2659391..2660779 [-], 1389 | putative pilus assembly protein | TraH_F, T4SS component |
| 17 | PMI2433 | 2660782..2661726 [-], 945 | putative pilus assembly | TraF_F, T4SS component |
| 18 | PMI2434 | 2662014..2663906 [-], 1893 | putative plasmid-related helicase | |
| 19 | PMI2435 | 2663903..2665867 [-], 1965 | putative plasmid-related protein | |
| 20 | PMI2436 | 2666041..2666241 [-], 201 | putative plasmid-related protein | |
| 21 | PMI2437 | 2666375..2667082 [-], 708 | putative plasmid-related protein | |
| 22 | PMI2438 | 2667172..2668245 [-], 1074 | putative plasmid-related protein | |
| 23 | PMI2439 | 2668337..2668678 [-], 342 | putative plasmid-related protein | |
| 24 | PMI2440 | 2668678..2669175 [-], 498 | putative plasmid-related DNA repair protein | |
| 25 | PMI2441 | 2669258..2670913 [-], 1656 | putative plasmid-related protein | |
| 26 | PMI2442 | 2670983..2671423 [-], 441 | putative plasmid-related protein | |
| 27 | PMI2443 | 2671485..2672438 [-], 954 | putative plasmid-related protein | |
| 28 | PMI2444 | 2672537..2673304 [-], 768 | putative plasmid-related protein | |
| 29 | PMI2445 | 2673304..2674263 [-], 960 | putative plasmid-related ATPase | |
| 30 | PMI2446 | 2674473..2675489 [-], 1017 | putative plasmid-related protein | |
| 31 | PMI2447 | 2675550..2675693 [-], 144 | putative plasmid-related protein | |
| 32 | PMI2448 | 2675777..2676595 [-], 819 | putative phage recombination protein | |
| 33 | PMI2449 | 2676675..2677094 [-], 420 | putative plasmid-related single-stranded DNA-binding protein | |
| 34 | PMI2450 | 2677110..2677436 [-], 327 | putative plasmid-related protein | |
| 35 | PMI2451 | 2677804..2678406 [+], 603 | putative plasmid-related protein | |
| 36 | PMI2452 | 2678515..2679189 [+], 675 | endonuclease precursor | |
| 37 | PMI2453 | 2679254..2679772 [-], 519 | putative plasmid-related regulatory protein | |
| 38 | PMI2454 | 2679792..2682065 [-], 2274 | putative plasmid-related membrane protein | |
| 39 | PMI2455 | 2682075..2682323 [-], 249 | putative plasmid-related glycin-rich membrane protein | |
| 40 | PMI2456 | 2682565..2686257 [-], 3693 | putative mating pair stabilization protein | TraN_F, T4SS component |
| 41 | PMI2457 | 2686260..2687288 [-], 1029 | putative plasmid pilus assembly | TraU_F, T4SS component |
| 42 | PMI2458 | 2687272..2688399 [-], 1128 | putative plasmid pilus assembly protein | TraW_F, T4SS component |
| 43 | PMI2459 | 2688407..2688919 [-], 513 | putative plasmid conjugation signal peptidase | TraF, T4SS component |
| 44 | PMI2460 | 2688903..2689250 [-], 348 | putative plasmid-related protein | |
| 45 | PMI2461 | 2689243..2691642 [-], 2400 | putative plasmid pilus assembly protein | TraC_F, T4SS component |
| 46 | PMI2462 | 2691642..2692334 [-], 693 | putative plasmid-related disulfide bond isomerase | TrbB_I, T4SS component |
| 47 | PMI2463 | 2692466..2693404 [-], 939 | putative plasmid-related protein | |
| 48 | PMI2464 | 2693397..2694230 [-], 834 | putative plasmid-related protein | |
| 49 | PMI2465 | 2694408..2694794 [-], 387 | putative plasmid-related protein | TraA_F, T4SS component |
| 50 | PMI2466 | 2694791..2695441 [-], 651 | putative plasmid-related protein | TraV_F, T4SS component |
| 51 | PMI2467 | 2695438..2696727 [-], 1290 | putative plasmid pilus assembly protein | TraB_F, T4SS component |
| 52 | PMI2468 | 2696730..2697629 [-], 900 | putative plasmid-related protein | TraK_F, T4SS component |
| 53 | PMI2469 | 2697610..2698236 [-], 627 | putative plasmid pilus assembly protein | TraE_F, T4SS component |
| 54 | PMI2470 | 2698233..2698514 [-], 282 | putative plasmid pilus assembly protein | TraL_F, T4SS component |
| 55 | PMI2471 | 2698550..2698789 [+], 240 | putative plasmid-related protein | |
| 56 | PMI2472 | 2698803..2699390 [+], 588 | putative plasmid-related protein | |
| 57 | PMI2473 | 2699417..2700061 [-], 645 | putative plasmid-related protein | |
| 58 | PMI2474 | 2700039..2700599 [-], 561 | putative plasmid-related protein | |
| 59 | PMI2475 | 2700609..2702429 [-], 1821 | putative plasmid conjugative transfer protein | TraD_F, T4SS component |
| 60 | PMI2476 | 2702478..2704628 [-], 2151 | putative plasmid conjugative relaxase | Relaxase, MOBH Family |
| 61 | PMI2477 | 2704877..2707012 [-], 2136 | putative DNA helicase | TraI_F, T4SS component |
| 62 | PMI2478 | 2707018..2713440 [-], 6423 | putative DNA helicase | |
| 63 | PMI2479 | 2713633..2718603 [-], 4971 | putative plasmid-related protein | |
| 64 | PMI2480 | 2718606..2721569 [-], 2964 | putative plasmid ATP-dependent helicase | |
| 65 | PMI2481 | 2721596..2722519 [-], 924 | putative plasmid-related protein | |
| 66 | PMI2482 | 2722837..2723136 [-], 300 | putative plasmid-related protein | |
| 67 | PMI2483 | 2723224..2724129 [-], 906 | putative plasmid-related exonuclease | |
| 68 | umuD | 2724786..2725235 [+], 450 | DNA repair protein | |
| 69 | umuC | 2725243..2726511 [+], 1269 | DNA repair protein | |
| 70 | PMI2485A | 2726550..2726720 [+], 171 | putative plasmid-related protein | |
| 71 | PMI2486 | 2726717..2726878 [+], 162 | conjugative transposon protein | |
| 72 | PMI2487 | 2727158..2727325 [+], 168 | putative conjugative transposon protein | |
| 73 | PMI2488 | 2727430..2728404 [+], 975 | putative plasmid-related protein | |
| 74 | PMI2489 | 2728407..2728676 [+], 270 | putative plasmid-related protein | |
| 75 | PMI2490 | 2728678..2729919 [+], 1242 | integrase | Integrase |
| 76 | PMI2491 | 2729936..2730130 [-], 195 | putative DNA-binding protein | |
| 77 | PMI2491A | 2730733..2730804 [-], 72 | N-terminus of peptide chain release factor 3 (fragment) | |
| 78 | PMI2492 | 2731174..2731710 [+], 537 | conserved hypothetical protein | |
| 79 | PMI2493 | 2731700..2732620 [+], 921 | conserved hypothetical protein | |
| 80 | rimI | 2732932..2733378 [-], 447 | ribosomal-protein-alanine acetyltransferase | |
| 81 | holD | 2733347..2733757 [-], 411 | DNA polymerase III, psi subunit | |
| 82 | rsmC | 2733887..2734900 [+], 1014 | ribosomal RNA small subunit methyltransferase C | |

