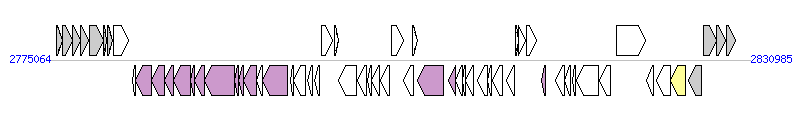The graph information of ICERsoGMI1000-1 components from AL646052 |
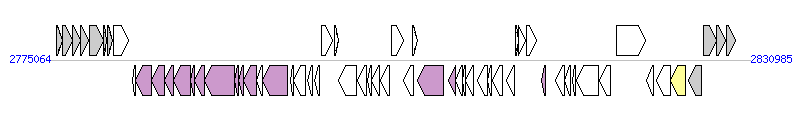 |
| Complete gene list of ICERsoGMI1000-1 from AL646052 |
| # | Gene  | Coordinates [+/-], size (bp) | Product
(GenBank annotation) | *Reannotation  |
| 1 | RSc2566 | 2775576..2776091 [+], 516 | putative methylated-dna--protein-cysteine methyltransferase | |
| 2 | RSc2567 | 2776088..2776840 [+], 753 | conserved hypothetical protein | |
| 3 | alkB | 2776851..2777507 [+], 657 | probable alkylated dna repair protein | |
| 4 | alkA | 2777509..2778147 [+], 639 | probable dna-3-methyladenine glycosylase protein | |
| 5 | ada | 2778230..2779312 [+], 1083 | probable ada regulatory of adaptative response contains: methylated-dna--protein-cysteine methyltransferase ec 2.1.1.63 o-6-methylguanine-dna transcription regulator | |
| 6 | RSc2571 | 2779312..2779581 [+], 270 | conserved hypothetical protein | |
| 7 | RSc2572 | 2779621..2780043 [+], 423 | probable lipoprotein | |
| 8 | tISRso15 | 2780153..2781373 [+], 1221 | isrso15-transposase protein | |
| 9 | RSc2574 | 2781673..2781915 [-], 243 | conserved hypothetical protein | |
| 10 | trbI | 2781912..2783189 [-], 1278 | probable conjugal transfer trbi transmembrane protein | TrbI, T4SS component |
| 11 | trbG | 2783192..2784196 [-], 1005 | probable conjugal transfer lipoprotein trbg | TrbG, T4SS component |
| 12 | trbF | 2784193..2784897 [-], 705 | probable conjugal transfer trbf transmembrane protein | TrbF, T4SS component |
| 13 | trbL | 2784928..2786301 [-], 1374 | putative conjugal transfer trbl transmembrane protein | TrbL, T4SS component |
| 14 | RSc2579 | 2786298..2786624 [-], 327 | putative lipoprotein | TrbJ, T4SS component |
| 15 | trbJ | 2786637..2787374 [-], 738 | probable conjugal transfer trbj signal peptide protein | TrbJ, T4SS component |
| 16 | trbE | 2787371..2789821 [-], 2451 | probable conjugal transfer protein trbe | TrbE, T4SS component |
| 17 | trbD | 2789834..2790106 [-], 273 | probable conjugal transfer trbd transmembrane protein | TrbD, T4SS component |
| 18 | trbC | 2790103..2790495 [-], 393 | probable conjugal transfer trbc transmembrane protein | TrbC, T4SS component |
| 19 | trbB | 2790492..2791559 [-], 1068 | probable conjugal transfer protein trbb | TrbB, T4SS component |
| 20 | RSc2585 | 2791556..2792032 [-], 477 | hypothetical ribbon-helix-helix harboring protein | |
| 21 | traG | 2792029..2794056 [-], 2028 | probable plasmid conjugation trag transmembrane protein | VirD4, T4SS component |
| 22 | RSc2587 | 2794292..2794555 [-], 264 | putative lipoprotein | |
| 23 | traR | 2794552..2795496 [-], 945 | probable plasmid replication regulatory trar transcription regulator protein | |
| 24 | limA | 2795654..2796046 [-], 393 | probable limonene-1,2-epoxide hydrolase protein | |
| 25 | limA2 | 2796232..2796618 [-], 387 | putative limonene-1,2-epoxide hydrolase protein | |
| 26 | RSc2591 | 2796769..2797659 [+], 891 | probable transcriptional regulator transcription regulator protein | |
| 27 | RSc2592 | 2797761..2798117 [+], 357 | hypothetical protein of unknown function duf419 | |
| 28 | RSc2593 | 2798145..2799572 [-], 1428 | putative carboxylesterase, type b; protein | |
| 29 | gstJ | 2799637..2800311 [-], 675 | probable glutathione-s-transferase protein | |
| 30 | RSc2595 | 2800406..2800735 [-], 330 | conserved hypothetical protein with dimeric alpha-beta barrel | |
| 31 | RSc2596 | 2800779..2801369 [-], 591 | putative amidase related to nicotinamidase; protein | |
| 32 | RSc2597 | 2801443..2802201 [-], 759 | putative short-chain dehydrogenase/reductase sdr; oxidoreductase protein | |
| 33 | RSc2598 | 2802374..2803273 [+], 900 | probable transcription regulator protein | |
| 34 | RSc2599 | 2803326..2804081 [-], 756 | hypothetical protein | |
| 35 | RSc2600 | 2804006..2804443 [+], 438 | putative substrate-binding domain related to lysr regulator protein | |
| 36 | RSc2601 | 2804432..2806483 [-], 2052 | putative type iv secretory pathway vird2 (relaxase)-related protein | Relaxase, MOBL Family |
| 37 | traF | 2806866..2807465 [-], 600 | probable conjugal transfer traf transmembrane protein | TraF, T4SS component |
| 38 | RSc2603 | 2807462..2808007 [-], 546 | conserved hypothetical protein | |
| 39 | RSc2604 | 2808004..2808288 [-], 285 | hypothetical ribbon-helix-helix protein | |
| 40 | parA3 | 2808276..2808914 [-], 639 | putative partition protein | |
| 41 | RSc2606 | 2809180..2810025 [-], 846 | probable replication initiation repa-like protein | |
| 42 | RSc2607 | 2810052..2810327 [-], 276 | putative dna binding protein | |
| 43 | RSc2608 | 2810442..2811212 [-], 771 | putative aspartate decarboxylase-like fold harboring protein | |
| 44 | RSc2609 | 2811557..2812150 [-], 594 | probable lipoprotein | |
| 45 | RSc2610 | 2812223..2812513 [+], 291 | putative transcription regulator protein | |
| 46 | RSc2611 | 2812522..2813085 [+], 564 | hypothetical tir protein | |
| 47 | RSc2612 | 2813107..2813955 [+], 849 | hypothetical transmembrane protein | |
| 48 | RSc2613 | 2814305..2814619 [-], 315 | hypothetical protein of unknown function duf736 | TraF, T4SS component |
| 49 | RSc2614 | 2815483..2816184 [-], 702 | conserved hypothetical protein | |
| 50 | RSc2615 | 2816196..2816681 [-], 486 | conserved hypothetical protein | |
| 51 | RSc2616 | 2816842..2817048 [-], 207 | conserved hypothetical protein | |
| 52 | RSc2617 | 2817109..2818902 [-], 1794 | conserved hypothetical protein | |
| 53 | RSc2618 | 2818983..2819813 [-], 831 | putative f plasmid gene 32-like protein | |
| 54 | RSc2619 | 2820359..2822662 [+], 2304 | conserved hypothetical protein | |
| 55 | radC2 | 2822729..2823238 [-], 510 | probable dna repair radc-like protein | |
| 56 | RSc2621 | 2823536..2824603 [-], 1068 | hypothetical protein of unknown function duf1016 | |
| 57 | int | 2824600..2825802 [-], 1203 | integrase-tyrosine recombinase protein | Integrase |
| 58 | purM | 2826032..2827093 [-], 1062 | probable phosphoribosylformylglycinamidine cyclo-ligase (airs)(phosphoribosyl-aminoimidazole synthetase) (air synthase) protein | |
| 59 | RSc2624 | 2827245..2828315 [+], 1071 | probable permease transmembrane transport protein | |
| 60 | RSc2625 | 2828338..2829039 [+], 702 | putative cog0593, atpase involved in dna replication initiation protein | |
| 61 | RSc2626 | 2829146..2829820 [+], 675 | probable had-superfamily subfamily ib, pspase-like; protein | |