The graph information of ICESpnCGSP14-1 components from CP001033 |
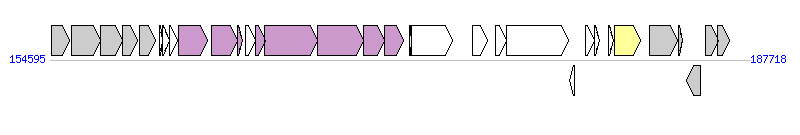 |
| Complete gene list of ICESpnCGSP14-1 from CP001033 |
| # | Gene  | Coordinates [+/-], size (bp) | Product
(GenBank annotation) | *Reannotation  |
| 1 | SPCG_0153 | 154645..155499 [+], 855 | lipoprotein | |
| 2 | SPCG_0154 | 155596..156969 [+], 1374 | hypothetical protein | |
| 3 | SPCG_0155 | 156962..158023 [+], 1062 | ABC transporter, ATP-binding protein | |
| 4 | SPCG_0156 | 158025..158717 [+], 693 | ABC transporter, permease protein, putative | |
| 5 | SPCG_0157 | 158833..159606 [+], 774 | transcriptional activator, Rgg/GadR/MutR family protein, C-terminal domain | |
| 6 | SPCG_0158 | 159788..159907 [+], 120 | hypothetical protein | |
| 7 | SPCG_0159 | 159930..160244 [+], 315 | hypothetical protein | |
| 8 | SPCG_0160 | 160260..160646 [+], 387 | hypothetical protein | |
| 9 | SPCG_0161 | 160675..162060 [+], 1386 | hypothetical protein | Orf21_Tn, T4SS component |
| 10 | SPCG_0162 | 162238..163443 [+], 1206 | Tn916, transcriptional regulator, putative | Relaxase, MOBT Family |
| 11 | SPCG_0163 | 163486..163707 [+], 222 | hypothetical protein | Orf19_Tn, T4SS component |
| 12 | SPCG_0164 | 163824..164321 [+], 498 | hypothetical protein | |
| 13 | SPCG_0165 | 164296..164802 [+], 507 | hypothetical protein | Orf17_Tn, T4SS component |
| 14 | SPCG_0166 | 164735..167233 [+], 2499 | hypothetical protein | Orf16_Tn, T4SS component |
| 15 | SPCG_0167 | 167236..169413 [+], 2178 | hypothetical protein | Orf15_Tn, T4SS component |
| 16 | SPCG_0168 | 169410..170411 [+], 1002 | hypothetical protein | Orf14_Tn, T4SS component |
| 17 | SPCG_0169 | 170408..171343 [+], 936 | hypothetical protein | Orf13_Tn, T4SS component |
| 18 | SPCG_0170 | 171618..171704 [+], 87 | tet(M) leader peptide | |
| 19 | SPCG_0171 | 171705..173639 [+], 1935 | tetracycline resistance protein TetM | |
| 20 | SPCG_0172 | 174564..175301 [+], 738 | rRNA methylase | |
| 21 | SPCG_0173 | 175656..176210 [+], 555 | resolvase | |
| 22 | SPCG_0174 | 176214..179132 [+], 2919 | transposase | |
| 23 | SPCG_0175 | 179188..179427 [-], 240 | hypothetical protein | |
| 24 | SPCG_0176 | 179923..180354 [+], 432 | hypothetical protein | |
| 25 | SPCG_0177 | 180351..180581 [+], 231 | hypothetical protein | |
| 26 | SPCG_0178 | 181042..181245 [+], 204 | hypothetical protein | |
| 27 | SPCG_0179 | 181327..182544 [+], 1218 | hypothetical protein | Integrase |
| 28 | SPCG_0180 | 182961..184277 [+], 1317 | hypothetical protein | |
| 29 | SPCG_0181 | 184315..184518 [+], 204 | hypothetical protein | |
| 30 | SPCG_0182 | 184713..185375 [-], 663 | hypothetical protein | |
| 31 | SPCG_0183 | 185632..186213 [+], 582 | hypothetical protein | |
| 32 | SPCG_0184 | 186159..186761 [+], 603 | hypothetical protein | |
