
| ICEfinder: a web-based tool for the detection of ICEs/IMEs of bacterial genomes |
| 1. Input data | |
|
1.1 You can upload one bacterial genome sequence and annotation in the GenBank format. [ 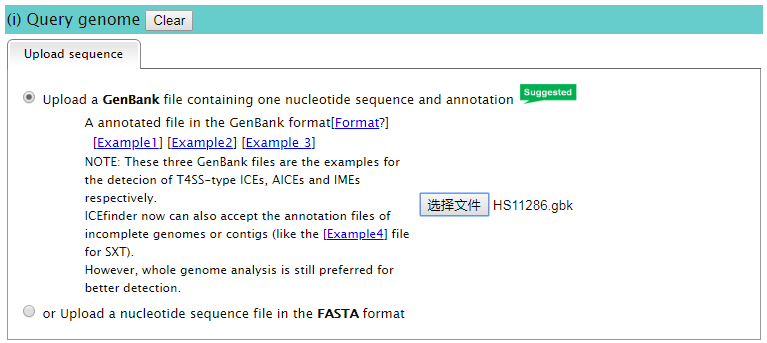
Note: When you download the GenBank file from the NCBI database, please select the GenBank(full) format, just as the following screenshot.  1.2 You can also upload one bacterial genome sequence in the FASTA format (the multi-FASTA file is acceptable). [ 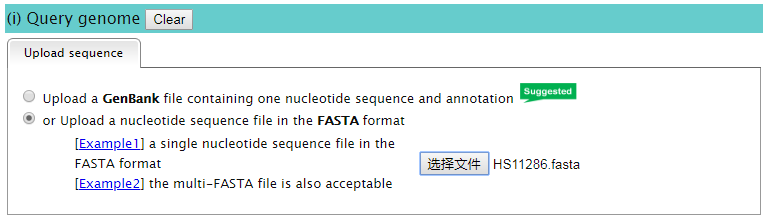 | |
| 2. Run ICEfinder - Predict | |
Get your input ready? ICEfinder will start to detect ICEs/IMEs immediately after you click the button “Run”.  Note: Depending on your input, the time consuming of ICEfinder may vary. When ICEfinder is running, the following page will be shown to users. 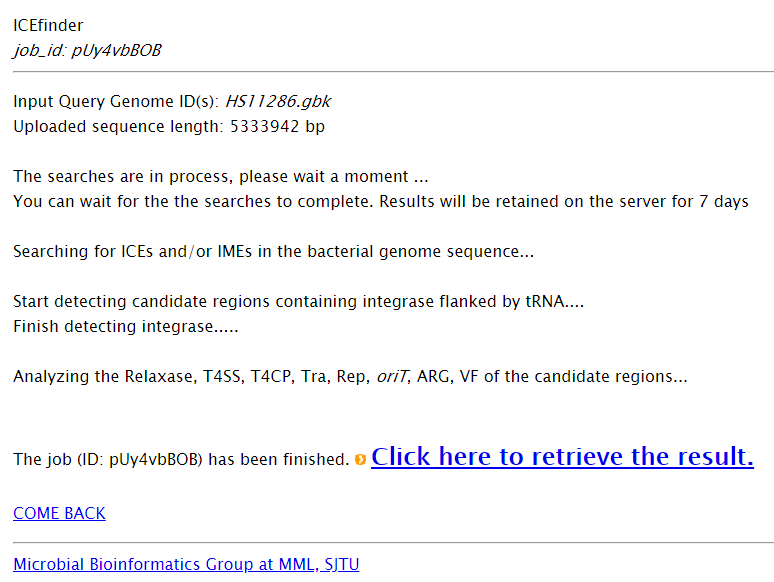 | |
| 3. Retrieve results - Predict | |
| You can get the analysis results with the Job ID or the link (shown in blue) provided on the running page.
Note: Please note that there is NO SPACE in your job ID provided by ICEfinder. As the Job ID is generated randomly, and may be not so easy to remember, we recommend that you just copy it to your clipboard, and paste when you retrieve the result. Sorry for the inconvenience it may cause.  |
|
| 4. Report of job | |
| Your analysis result is shown on the report page. All the predicted ICEs/IMEs in the bacterial genome are listed in the summary tab.
If you need further detail of each ICE/IME, please click on the link shown in blue.  | |
|
Note: You can also click the above image to have a larger view. The user can find all the related information towards each ICE/IME predicted by ICEfinder,which is shown under four parts:the "Summary" part, the "Map" part, the "Gene List" part, and the "Download" part. Moreover, we also provide a ICEfinder_compare tool to compare the predicted ICE/IME with the ones collected in ICEberg2 database. | |
"Summary" part looks like this: "Map" part looks like this: 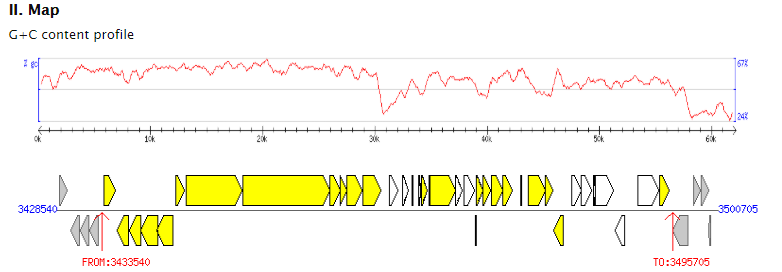 "Gene List" part looks like this: 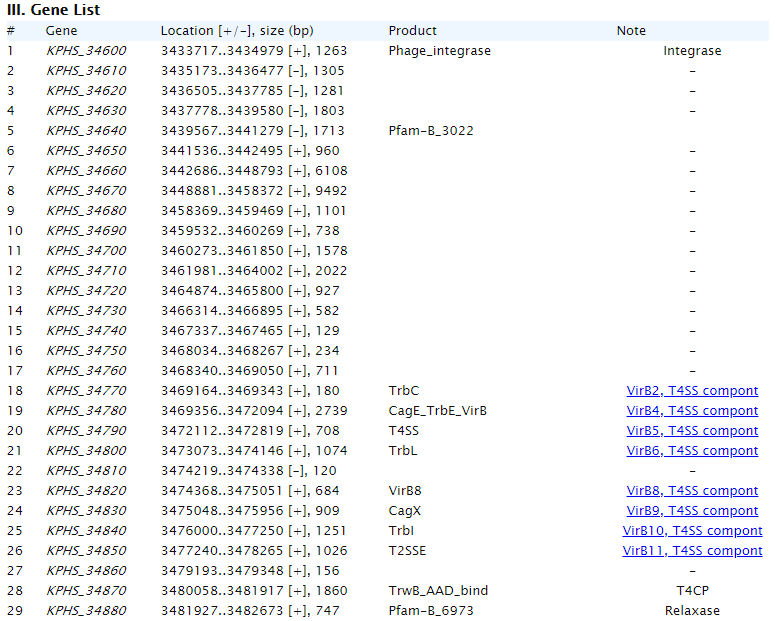 "Download" part looks like this:  "Comparison" part looks like this: 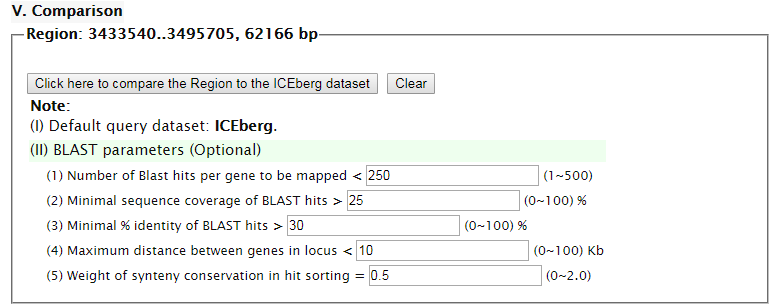 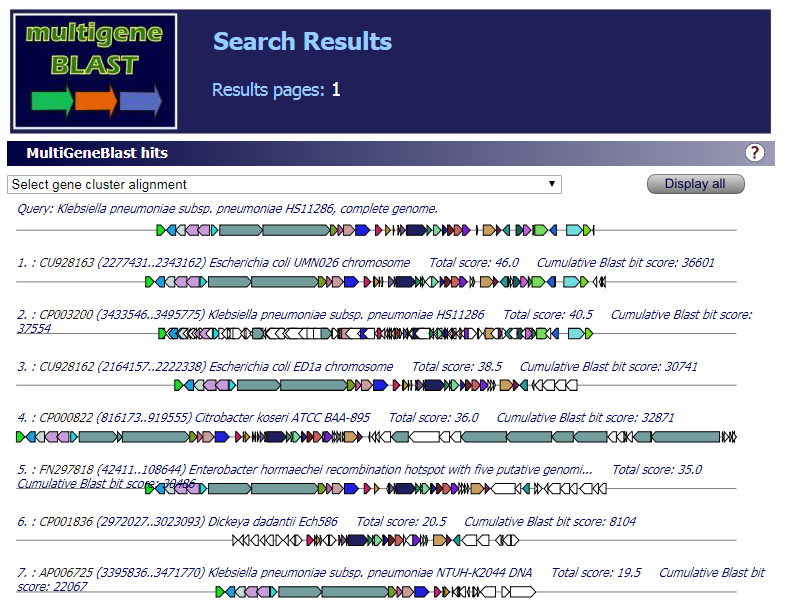 | |